Polyploid speciation
(Mal)adaptive consequences
of inter-ploidy gene flow
Genome duplication (polyploidisation) is a dominant force in sympatric speciation, particularly in plants and including many crops. It is usually assumed that polyploidy poses an instant barrier to gene flow between diploid and its polyploid derivative. Our genomic survey, however, demonstrated rampant inter-ploidy gene flow in wild Arabidopsis. Yet mechanisms allowing such genome permeability and its evolutionary consequences remain elusive. In this project, we aim to dissect drivers and mechanisms of ploidy barrier in five plant species and test the hypothesis that selection may promote inter-ploidy introgression.
Firstly, we will infer mechanisms shaping inter-ploidy barrier by crossing experiments and transcriptomics. Specifically, continue further with addressing genetic mechanisms underlying triploid block and its ecological consequences in natural populations of several mixed-ploidy plant systems such as Arabidopsis arenosa, Cardamine amara, Mimulus guttatus, Medicago falcata and Dactylis glomerata
We pioneered these questions in Arabidopsis arenosa where we firstly detected rampant gene flow across ploidy repeatedly in three contact zones. We thus followed up by field surveys and crossing experiments to infer the strength and nature of such imperfect barrier. Using field sampling and flow cytometry of mixed-ploidy (2x+4x) populations we documented a lack of triploid (3x) adult hybrids among ~4500 cytotyped plants and two 3x seedlings germinated from in situ collected seeds. This is not due ecological separation as we found no distinction in local habitat preferences among the ploidies.
Experimental crossings
To test whether such lack of hybrids is due to intrinsic post-pollination barriers we performed experimental crossings. The inter-ploidy crossing progeny was less fertile and majority of germinated seedlings were triploids, although we also found a non-negligible proportion of tetraploid hybrids (~7 %), documenting the possible role of unreduced gametes in mediating gene flow.
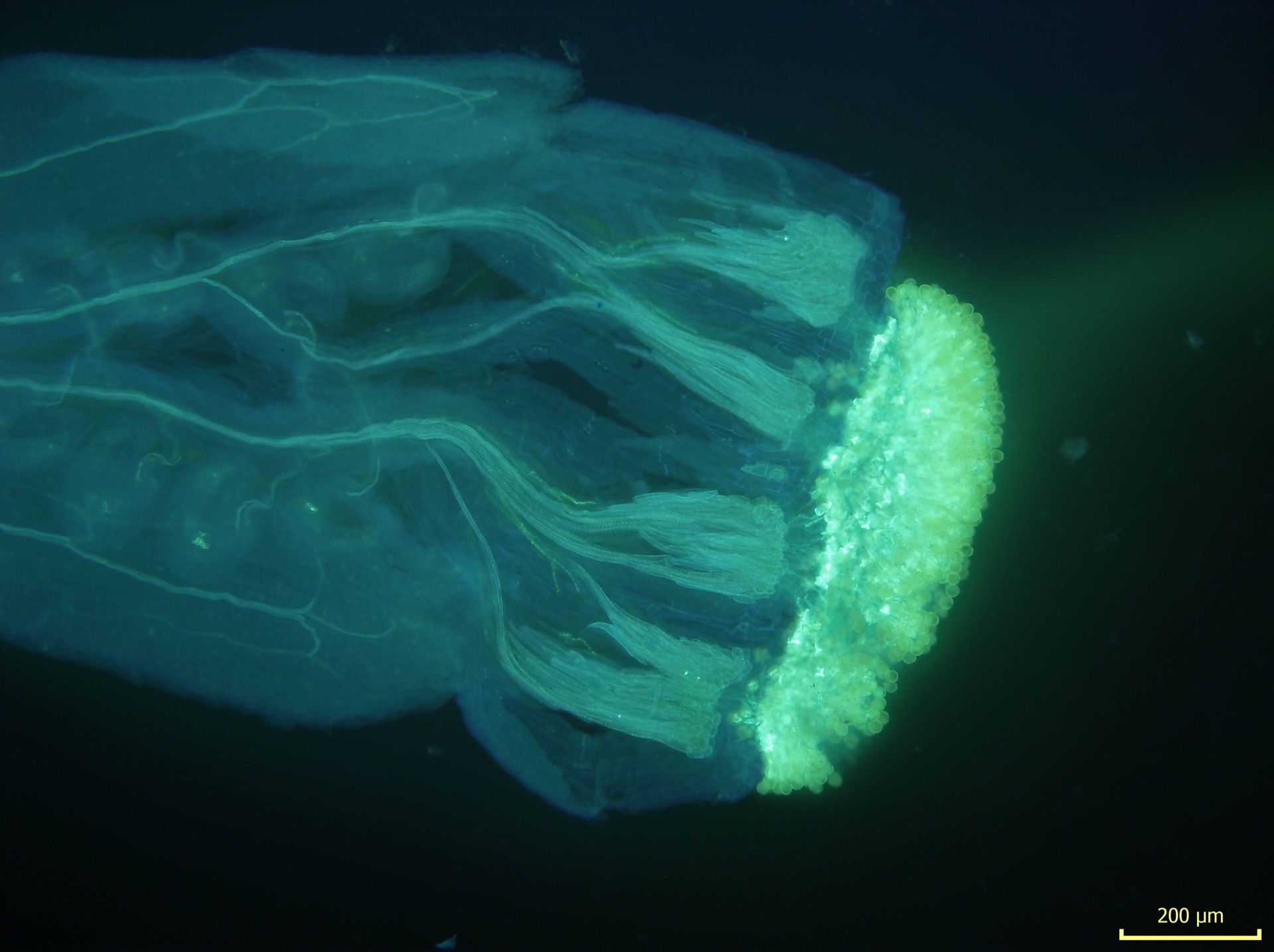
We used confocal microscopy and observed clear endosperm developmental defects in reciprocal interploidy crosses. These results document strong yet still incomplete triploid block in A. arenosa. The complete absence of adult triploid hybrids in the field suggests alternative mechanisms of cross-ploidy gene flow such as involvement of unreduced gametes and formation of one-step tetraploid hybrids.

Finally, we will infer genomic regions susceptible vs. resistant to interploidy introgression. The results have the potential to shift our perception of polyploidy towards speciation-with-gene-flow scenaria and inform breeding programmes involving polyploid crops.


